 
Therefore, lysosomes are a target for a number of therapeutics and there are several methods available to study the intracellular fate and localization of NP in lysosomes. Lysosome dysfunction is associated with multiple diseases, such as cancer, inflammatory and neurodegenerative diseases, and Tay Sachs disease. Lysosomes are dynamic cellular catabolic organelles responsible for the degradation of macromolecules and play a key role in the accumulation of internalized materials such as NP. Finally, depending on their properties, NP localize in the subcellular compartments, including lysosomes, endoplasmic reticulum, Golgi apparatus, mitochondria and nucleus or bypass these compartments and end up in the cytoplasm. Upon deposition at the outer cell membrane, the NP are internalized via a variety of endocytic mechanisms and pass through an endosomal sorting process. The use of engineered nanoparticles (NP) for biomedical applications requires a detailed understanding of the NP-cell interaction, NP uptake and intracellular fate. Continuous live cell imaging of J774A.1 cells. Comparison of raw integrated intensity between live and fixed cells. Comparison of Pearson’s and Manders’ coefficients between live and fixed cells. Representative images of LysoTracker Red probe and Lamp-2 stainings in a single cell. Representative bright field image of live cells (A) and fluorescent microscopy image of fixed cells (B) showing a population of individual cells used for colocalization analysis. Representative images of individual cells from different experiments for identification and contour with the corresponding histogram of the complete image acquired for each individual channel (nanoparticles and lysosomes). Size distributions of 119 nm SiO 2-Cy5 particles and 920 nm SiO 2-Cy5 particles measured by dynamic light scattering. 
Size distributions of 59 nm SiO 2-BDP FL NP and 59 nm SiO 2-RhoB NP measured by dynamic light scattering. Transmission electron micrographs (TEM) of the different silica (nano)particles. 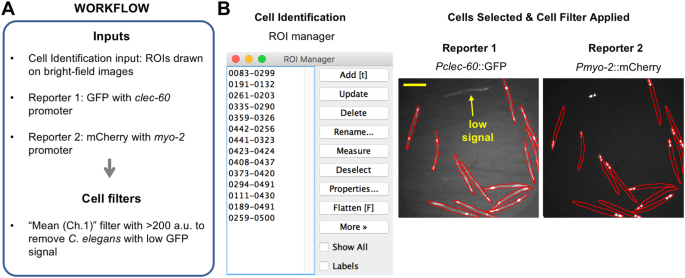
Confocal microscopy images of different LysoTracker probes. 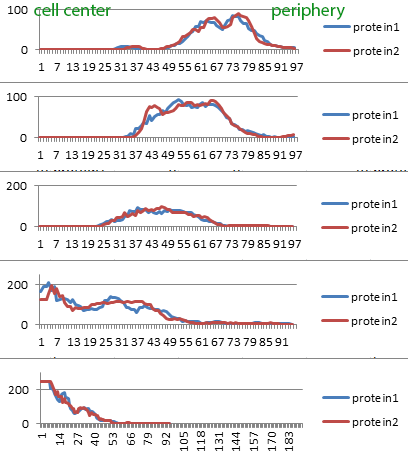
Comparison of the absorbance spectra of the different (nano)particles. Comparison of the emission spectra of the different (nano)particles. Comparison of the absorbance and emission spectra of the different LysoTracker probes. The Creative Commons Public Domain Dedication waiver ( ) applies to the data made available in this article, unless otherwise stated in a credit line to the data.Īdditional file 1: Figure S1A. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. I found a good cell counting tool for ImageJ lacking, so maybe this plugin (and the other) can be included as a standard part of FIJI.Open AccessThis article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The ImageJ plugin 1 might be somewhat hard to understand how to use effectively (though we hope not), but ImageJ plugin 2 should be very simple and useful to the broader community. I hope the community will appreciate our work. *3D visualize cells according to colocalization data *Export combined image series data to Matlab for 3D modeling *Analyze and edit data from image series. *Import data from Colocalization Object Counter *Save data, load data, and export data to Excel. *Tools for subsequent 3D modeling/representation of data: draw tissue contours and indicate image-series global XY-origin. *Assign, classify and keep track of multichannel signal presence/absence (colocalization analysis) for each cell/object. *Quantity (count) cells/objects in a semi-automatic manner. ImageJ plugin 2: Colocalization Object Counter: This simplifies the analysis of 3D colocalization data. *Can transform Z-stack 3D data into a specialized 2D Z-projection where Z-projection colocalization artifacts are removed/reduced. *Designed to help avoid common colocalization analysis artifacts and errors. *Pre-process multichannel Z-stack (or 2D) microscopy images into a visual format for faster, simpler, and more accurate colocalization analysis. ImageJ plugin 1: Colocalization Image Creator: FIJI update site (add to install plugins): Index of /ObjectColocalizationPlugins 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
